MENU
The Electronic Scholarly Publishing Project: Providing world-wide, free access to classic scientific papers and other scholarly materials, since 1993.
More About: ESP | OUR CONTENT | THIS WEBSITE | WHAT'S NEW | WHAT'S HOT
Believe It or Else !
The following cartoon appeared in newpapers in March of 1997.
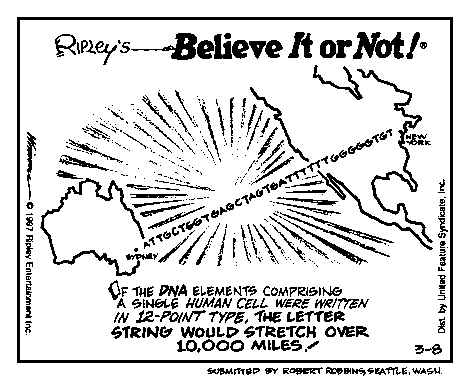
BELIEVE IT OR NOT, getting this single sentence written and approved took more drafts, more revisions, more consultation, and more editorial effort than that spent on many scientific papers.
The story of how this "major" publication came to be is detailed below...
A haploid copy of the human genome contains 3.3 billion base pairs of DNA.
Typing that out as a list of A's, C's, T's, and G's in 10-pitch font (i.e., 10 characters per inch) would, by definition, generate a string of letters over 5000 miles long, since
3,300,000,000 / 5280 / 12 / 10 = 5208
Over many years of teaching, I used this simple fact to help people understand the scale of genomic numbers.
Then, in October of 1996, I received the following unsolicited email message.
Date: Thu, 24 Oct 1996 16:07:02 -0400
From: editor@ripleys
To: rrobbins@gdb.org
Subject: Believe It or Not!
Robert:
I spoke to Steven Salzberg regarding scientific facts to feature in the
Ripley's Believe It or Not! cartoon. He cited a DNA illustration he uses in
teaching. Though I may not express this properly. He suggests that if all
the letter groups associated with one DNA helix were printed in 10 pt. type
the string would be thousands of miles long! He noted that you are the
author of this illustration and passed along your E-Mail address so that I
may speak with you regarding this illustration and any other facts such as
this which are typical of the kinds of things upon which Ripley's thrives.
Please E-Mail a phone # and convenient time (your zone) when we may speak.
Thanks!
I responded to this inquiry, noting that 3.3 billion divided by 5280 divided by 12 divided by 10 was indeed equal to 5208, but added that this result from simple division wasn't much of an insight.
The editor indicated that it was still of interest to Ripley's and asked if I would help draft a few words to summarize this factoid so that it could be used in a Believe It or Not! cartoon.
I said, "sure," and only later discovered that this would require an exchange of many emails and voice mails, over several months, as we tried to converge on something that was simultaneously accurate, simple, and brief.
Early discussions hinged on the difference between 10-pitch and 10-point font. Pitch is a measure of characters per inch, and point is a direct measure of size, where "point" is the unit of measure. It turns out that 10-pitch font is approximately 12 points in height, so we agreed to refer to "12-point font".
A more significant problem occurred, when distinguishing between diploid and haploid cells was deemed too obscure, and the proposed description:
Every human cell contains two sets of DNA, one received from each parent.
DNA is built as a string of four different components, so a DNA molecule
can be described as a long string of four different letters:
ATTCGGCTAC...
If we tried to write out just one set of human DNA as a string of letters
in 12-point type, then that string would stretch from .... to ....
was deemed simultaneously too verbose and too complex.
An effort to eliminate the tricky concept of "one set of human DNA" by referring to "the DNA in a single human sperm cell" proved too racy for appearance in a family newspaper.
Finally, after three months of kicking ideas around, the editor suggested we ignore the redundancy in a diploid cell and just double the length in miles:
Date: Thu, 9 Jan 1997 12:39 (EST)
From: editor@ripleys
To: rjr8222@gmail.com
Subject: Believe It or Not!
Bob:
Try this on for size........
Believe It or Not ! If the elements comprising the DNA in a single human
cell were written in 12 point type, the letter string would be over 10,000 mi
in length!
I would use a cartoon of a globe with two cities appropriate in distance from
one another connected by a ribbon of A.C.T.G.'s
I agreed immediately and thought the publication was finally done. But, a few weeks later, the editor contacted me, wanting to know what kinds of patterns of A's, T's, C's, and G's should or should not be used if the illustration was to look like "real DNA." I responded by sending him a copy of the full genomic sequence for human beta-hemoglobin, taken from GenBank.
This pleased Ripley's, and finally the deed was done, and the cartoon appeared in March.
Did they use part of the hemoglobin sequence?
Maybe yes, maybe no. A BLAST search of GenBank using the sequence in the cartoon shows the following:
HUMHBB U01317
(73308) Human beta globin region on chromosome 11.
Length = 73,308
Plus Strand HSPs:
Score = 97 (26.8 bits), Expect = 1.3, P = 0.92
Identities = 25/32 (78%), Positives = 25/32 (78%)
Query: 1 ATTGCTGGTGAGCTAGTGATTTTTTGGGGGTG 32
||||||||||||||||| |||| ||| |
Sbjct: 29756 ATTGCTGGTGAGCTAGTATGATTTTTTGGGGG 29787 |
The match is not perfect, but is too close to be due to chance. Apparently "real" DNA needed editing, too.
Despite all the above, the project was fun and I am looking forward to my next opportunity to publish a major finding in Ripley's Believe It or Not! — perhaps the one about synthesizing life from materials purchased in a drugstore, or maybe elucidating fully the physical substrate of mind.
Robert J. Robbins
rjr8222@gmail.com
ESP Quick Facts
ESP Origins
In the early 1990's, Robert Robbins was a faculty member at Johns Hopkins, where he directed the informatics core of GDB — the human gene-mapping database of the international human genome project. To share papers with colleagues around the world, he set up a small paper-sharing section on his personal web page. This small project evolved into The Electronic Scholarly Publishing Project.
ESP Support
In 1995, Robbins became the VP/IT of the Fred Hutchinson Cancer Research Center in Seattle, WA. Soon after arriving in Seattle, Robbins secured funding, through the ELSI component of the US Human Genome Project, to create the original ESP.ORG web site, with the formal goal of providing free, world-wide access to the literature of classical genetics.
ESP Rationale
Although the methods of molecular biology can seem almost magical to the uninitiated, the original techniques of classical genetics are readily appreciated by one and all: cross individuals that differ in some inherited trait, collect all of the progeny, score their attributes, and propose mechanisms to explain the patterns of inheritance observed.
ESP Goal
In reading the early works of classical genetics, one is drawn, almost inexorably, into ever more complex models, until molecular explanations begin to seem both necessary and natural. At that point, the tools for understanding genome research are at hand. Assisting readers reach this point was the original goal of The Electronic Scholarly Publishing Project.
ESP Usage
Usage of the site grew rapidly and has remained high. Faculty began to use the site for their assigned readings. Other on-line publishers, ranging from The New York Times to Nature referenced ESP materials in their own publications. Nobel laureates (e.g., Joshua Lederberg) regularly used the site and even wrote to suggest changes and improvements.
ESP Content
When the site began, no journals were making their early content available in digital format. As a result, ESP was obliged to digitize classic literature before it could be made available. For many important papers — such as Mendel's original paper or the first genetic map — ESP had to produce entirely new typeset versions of the works, if they were to be available in a high-quality format.
ESP Help
Early support from the DOE component of the Human Genome Project was critically important for getting the ESP project on a firm foundation. Since that funding ended (nearly 20 years ago), the project has been operated as a purely volunteer effort. Anyone wishing to assist in these efforts should send an email to Robbins.
ESP Plans
With the development of methods for adding typeset side notes to PDF files, the ESP project now plans to add annotated versions of some classical papers to its holdings. We also plan to add new reference and pedagogical material. We have already started providing regularly updated, comprehensive bibliographies to the ESP.ORG site.
ESP Picks from Around the Web (updated 28 JUL 2024 )
Old Science

Weird Science

Treating Disease with Fecal Transplantation
Fossils of miniature humans (hobbits) discovered in Indonesia
Paleontology

Dinosaur tail, complete with feathers, found preserved in amber.
Astronomy

Mysterious fast radio burst (FRB) detected in the distant universe.
Big Data & Informatics

Big Data: Buzzword or Big Deal?
Hacking the genome: Identifying anonymized human subjects using publicly available data.